2026 Conference
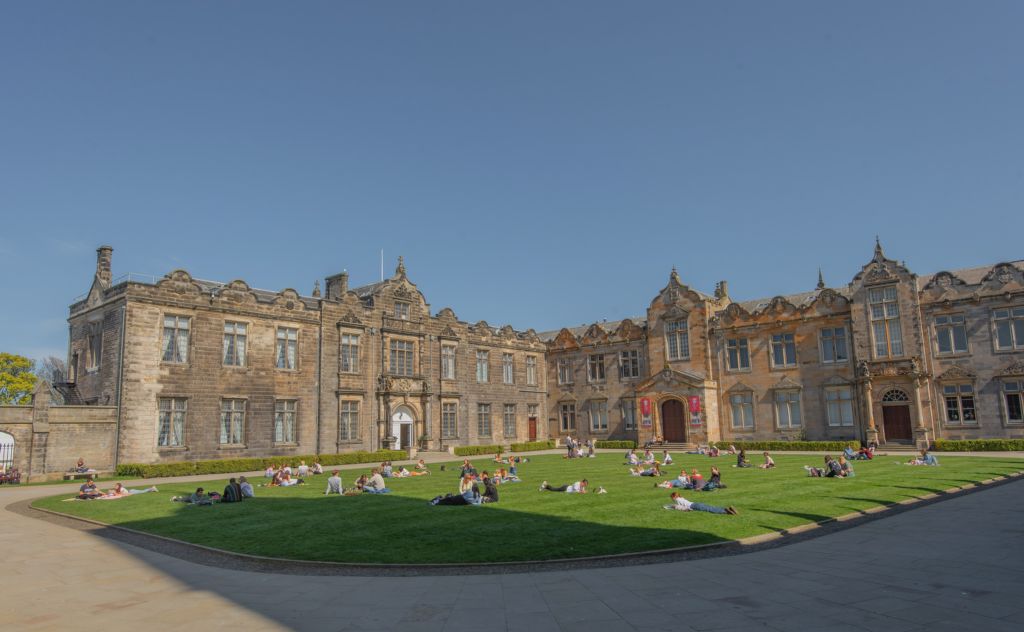
The University of St Andrews looks forward to welcoming the fifth BioInference meeting on 10th – 12th June 2026!
Participants can look forward to a welcoming atmosphere, lively academic conversations, and social events set against beautiful coastal scenery. The University of St Andrews—Scotland’s oldest university, founded in 1413—is nestled in a charming seaside town celebrated for its rich academic heritage, medieval architecture, and unique character. St Andrews is also the birthplace of golf and home to the world-famous Old Course, with a compact town centre that is easy and enjoyable to explore on foot.
Conference Information
We are excited to welcome you to a three-day conference featuring a range of stimulating presentations and networking opportunities. Below you can find the tentative timetable for the three days of the conference (subject to revisions).
📅 Wednesday, June 10
🌅 Morning
🕘 09:00–09:40 — Registration & Welcome 🎤 09:40–11:00 — Talks ☕ 11:00–11:30 — Coffee Break 🎤 11:30–13:00 — Talks
🍽️ Noon
🥗 13:00–14:00 — Lunch
🌇 Afternoon
🎤 14:00–15:00 — Talks ☕ 15:00–15:30 — Coffee Break 🪧💬 15:30–17:00 — Poster Session
🌙 Evening
🎉 Social Events ⛳ 🏰 🌊 🥃🍸
📅 Thursday, June 11
🌅 Morning
🎤 09:30–11:00 — Talks ☕ 11:00–11:30 — Coffee Break
🍽️ Noon
🏭 11:30–13:00 — Industrial Partners Session 🥗 13:00–14:00 — Lunch
🌇 Afternoon
🎤 14:00–15:30 — Talks ☕ 15:30–16:00 — Coffee Break 🎤 16:00–17:00 — Talks
🌙 Evening
🍷 Conference Dinner
📅 Friday, June 12
🌅 Morning
🎤 09:30–11:00 — Talks ☕ 11:00–11:30 — Coffee Break 🎤 11:30–13:00 — Talks 🏁 Wrap-up
🏆 13:00–13:15 — Awards & Closing Remarks 🧳 13:15–14:00 — Lunch & Departure
Social activities
Optional social activities will take place on Wednesday evening. These include opportunities to explore the local culture of St Andrews, its rich golf history, and the Scottish traditions of whisky and gin tasting.
Some of the optional social activities involve additional fees. You can find out more details and indicate your preferences on the practical information page.
Prizes
We are pleased to announce that this year four prizes will be awarded as part of the conference. Each one of the awards comes with a monetary prize of £200:
- Reproducibility prize for the individual whose research (presented at the conference) is judged to be the most straightforward to reproduce. This is judged based on submitted code. This prize is open to all conference presenters.
- Transparency in Research prize to promote clear and open presentation of research methods and encourage critical and honest assessment of research outcomes. This prize is also open to all conference presenters.
- Best Early Career Research talk and Best Early Career Research poster presentation, decided by the conference participants.
All contributing oral and poster presenters at the conference are invited to self-nominate themselves for the Transparency in Research and Reproducibility prizes by sending through the appropriate documentation via email to bioinference@gmail.com by May 29th.
The winners of the three prizes will be announced on Friday afternoon (12th of June) during the Closing remarks of the conference.
Transparency in Research
The new Transparency in Research prize aims to promote clear and open presentation of research methods, open and honest discussion of failures as well as successes in the path of research activities, and encourage critical and honest assessment of research outcomes.
As it is not possible to present all of the above during a talk or on a poster, to be considered for this prize we ask you to send a short paragraph (200-300 words) by email to bioinference@gmail.com with the subject title “Transparency in Research prize” by May 29th.
The following is guidance for the motivational paragraph (200-300 words) to be sent by email. The focus of the paragraph should be on the transparency of your work, on the lessons learned when developing your method, or applying your methodology, and the limitations of material, methods and results. Aim to answer at least one of the following questions in the paragraph:
- What are the take home messages you would want another researcher to know working on a similar topic? This could be a method that you have used and worked well or badly in your problem.
- Can you recommend one aspect of best practice for other researchers/ practitioners based on your experience in the development of the presented work?
- What are the aspects of your method that you would like to improve?
- Are there any other lessons you learnt during the process of developing your analysis and would like to share?
We acknowledge that the paragraph and especially the time of the presentation is not sufficient to share all the relevant information. Therefore we encourage you to focus your presentation on the most important aspects of your work and balance transparency alongside other aspects covered.
A prize committee will review both the submitted paragraphs as well as the presentations for deciding on the winner of this prize.
Reproducibility Award:
As we have done in previous years, we are allowing conference presenters to put themselves forward for a Reproducibility Award. To be considered for this award, please email us a link to a public repository (e.g. a GitHub repo) where the software is housed at bioinference@gmail.com with the subject title: “Reproducibility Award” by May 29th.
To be eligible for the award, entrants must satisfy the following criteria:
- The software must be coded in an open-source language
- The repository holding the software must be openly available
- The software must underpin work being presented at BioInference 2026
The work will be assessed according to the following criteria:
- How well the Readme file explains how to reproduce analyses
- Are software requirements (e.g. packages) well described?
- Are software requirements easy to acquire?
- Does the repository have descriptive (e.g. tutorial) examples?
- Does the repository have a license (e.g. an MIT License) which includes its permitted usage?
- Does the functionality have high quality documentation?
If you have any questions about these two prizes please do not hesitate to contact us at bioinference@gmail.com.
Travel costs funding
We are grateful for the financial support that we have secured from several sponsors that enables us to support travelling costs for ECRs (anyone who does not have a permanent academic position).
Similarly to previous BioInference editions and depending on demand, we may provide a maximum amount of £200 towards travelling costs. To be considered for travel support, please indicate this on the registration page (see related question) and keep all travel receipts. More information about submitting your travel expenses will be sent out after the conference.
We are grateful that a portion of this funding is from the Glasgow Mathematical Journal Learning and Research Support Fund that is designed to support travel costs for Scottish-based ECRs.
Additional travel support opportunities are also available, including the following:
- For members of the European Society for Mathematical and Theoretical Biology (ESMTB) through https://www.esmtb.org/Travel-Support. Applicants are allowed to become a member (for up to 25€ per year!) before applying for funding. Note that the application must be done BEFORE the conference, with a notification typically within one month after the submission. The deadline has now passed.
- For ECRs members (with at least one year of membership) of the Society for Mathematical Biology (SMB) through https://www.smb.org/travel-grants.
Featured Talks
A booklet containing the schedule for all the talks their abstracts, and other additional information is now available here. Please find below the list of all talk titles:
- Tarek Alrefae (University of Oxford) - Interpretable measures for sensitivity and identifiability in models of infectious disease
- Alejandra Avalos Pacheco (JKU Linz) - Rotational Invariant Sparse Factor Models with the l1-ball prior
- Feargus Ball (University of Birmingham) - Modelling the Spread of Acinetobacter baumannii in an Intensive Care Unit
- André Brito (NOVA Math Portugal) - Coupling Environmental Exposure and Human Mobility in an Agent-Based Model for Influenza Transmission
- Arianna Ceccarelli (University of Oxford) - A Bayesian inference framework to calibrate one-dimensional velocity-jump models for single-agent motion using discrete-time noisy data
- Yari Cerruti (University of St Andrews) - Modeling varying demography along phylogenies from large-scale genomic data
- Lloyd Chapman (Lancaster University) - Bayesian inference for individual-level epidemic models
- Junjie Chen (University of Oxford) - Understanding the Biology Underpinning the Interaction between Arboviruses and Aedes Aegypti Mosquitoes Using Mathematical and Statistical Modelling
- Claudia Chien (Charité-University Hospital Berlin) - Intra-individual longitudinal white matter tract integrity modelling from diffusion weighted images
- Diogo da Silva Ribeiro (University of St Andrews) - Inferring gene flow from phylogenies with too many genomes
- David Dreifuss (Imperial College London) - Statistical Methods for Wastewater-Based Genomic Epidemiology
- Anders Erlandson (University of Glasgow) - Quantifying the effects of praziquantel treatment on Schistosoma infection dynamics using imperfect diagnostics
- Sokratia Georgaka (University of Manchester) - Non-Negative Matrix Factorisation for Multimodal Analysis of Spatial Omics: From Shared Latent Factors to Proteomics-Constrained Cell-Type Mapping
- Andrew Golightly (Durham University) - Robust, partially alive particle Metropolis-Hastings for stochastic kinetic models
- Lorna Jamieson (University of St Andrews) - Applying Spatial Models to Opinion Dynamics
- Toby Kettlewell (University of Glasgow) - ZINBGT: Exploratory data analysis of transcriptomic expression using mixture models
- Jan-Ole Koslik (Bielefeld University) - Fast estimation of state-switching correlated random walk models subject to measurement error
- Rajneesh Kumar (University of Sussex) - Soil Biodiversity, Ecological Niches, and Climate Change: Modelling Insights from the BIOservicES Project
- Jochen Kursawe (University of St Andrews) - Data-driven modelling of spatial actomyosin dynamics during pulsed cellular contractions
- Theo Kypraios (University of Nottingham) - Neural Posterior Estimation for Stochastic Infectious Disease Models
- Xiahui Li (University of St Andrews) - An Entropy-based Approximate Bayesian Inference Method for Compartmental Models in Epidemiology
- Marta Belchior Lopes (Universidade NOVA de Lisboa) - Learning from high-dimensional omics data: sparse statistical and machine learning approaches in brain cancer
- Claus Mayer (Biomathematics & Statistics Scotland) - A p-value combination method for random effects meta-analysis of high-dimensional omics data
- Claire McWhite (University of Arizona) - A General Algorithm for Detecting Higher-Order Interactions via Random Sequential Additions
- Theodosios Papazoglou (University of Hong Kong) - Bayesian Neuronized Knockoff Supervised Principal Component Analysis
- Linus Schumacher (University of Edinburgh) - Inferring rules of tissue organisation with neural cellular automata
- Lucas Siu (Imperial College London) - Functional fine-mapping using variational inference
- Nikolaos Stefanidis (University of Sheffield) - Parameterising Notch-mediated cell state transitions in human enteric nervous system development
- Massimiliano Tamborrino (University of Warwick) - Network inference via approximate Bayesian computation. Illustration on a stochastic multipopulation neural mass model
- Frederick Truman-Williams (University of Oxford) - Simulating stochastic population dynamics: The Linear Noise Approximation can capture non-linear phenomena
- Sara Wade (University of Edinburgh) - Decoding the brain: Statistical learning for high-throughput neuroanatomy
- Hanyu Wang (University of St Andrews) - Divide and Conquer techniques for Bayesian mixture models with evaluation of uncertainty
- Michael Whitehouse (Imperial College London) - Scalable calibration for partially observed individual-based epidemic models through categorical approximations
- Konstantinos Zygalakis (University of Edinburgh) - Hybrid modelling for computation and parameter inference of spatial and non-spatial chemical kinetics
Featured Posters
A booklet containing the abstracts of all featured posters is now available here. Please find below the list of all poster titles:
- Begoña Bolos (University of Edinburgh) - The C-index Multiverse
- Sian Dennett (University of Oxford) - Understanding Spatiotemporal Variability in Dengue Transmission Dynamics in Brazil
- Marina Evangelou (Imperial College London) - Adaptive gene and pathway selection through spare-group SLOPE
- Chao Gao (University of St Andrews) - Decoding Hidden Signals: A Supervised Learning Framework for Information Estimation
- Alicia Gill (University of Oxford) - Estimating Lassa Fever Vaccine Efficacy Under Diagnostic Uncertainty
- Charis Hanna (University of St Andrews) - IoMB: Evaluating Object Detectors on Occluded and Imbalanced Seabird populations
- Grant Henderson (Biomathematics and Statistics Scotland) - Quantifying Spread and Control of UK HPAI Outbreaks
- Ishtiaq Hussain (University of Strathclyde) - Mathematical Modelling for Using Insect-Specific Viruses to Curb Dengue Epidemics: A Case Study in Pakistan
- Shilji Nihar Kadavath madathin padayantavida (University of Lancashire) - Country level variations in time-to-recurrence after curative resection of colorectal liver metastasis (CRLM) patients: A multinational retrospective cohort analysis
- Cameron Kerr (University of Edinburgh) - Decoding Clonal Dynamics in Blood Cancer Through An Adapted CHIP Model
- Richard Kettle (University of Edinburgh) - Statror Reveals Context Dependent Cytokine Responses in CD8+ T Cells
- Janique Krasnowska (University of Warwick) - Coalescence in Multi-Type Branching Processes with Applications to Dormancy
- Mohammed Moinuddin (University of Lancashire) & Halima Kabir Tonni (University of Lancashire) - Identifying the factors that moderate the association between resection margin and survival outcomes for colorectal liver metastasis patients in Europe
- Francesco Morandin (University of Pisa) - From Genes Correlation to Differential Expression: COTAN scRNA-seq Analysis
- Akihiro Mori (University of Aberdeen) - Hybrid stochastic modelling of protocell growth and homochirality emergence
- Scott Pirrie (Optima Partners Ltd) - Integrating genetics and multi omics to distinguish risk and progression targets for drug repurposing in Parkinson’s disease
- Andrey Shternshis (Uppsala University) - Learning dynamics of microbiome abundance by logit-normal processes
- Alexander Zarebski (University of Cambridge) - GENIE: A Spatio-Temporal Neural Network for Probabilistic Disease Forecasting using Agent-Based Model simulations
We are grateful for the financial support from:
- The University of St Andrews
- Elsevier
- European Society for Mathematical and Theoretical Biology (ESMTB)
- SMB. The Society will also consider applications for travel support
- The Edinburgh Mathematical Society
- The Glasgow Mathematical Journal Trust
Conference Organisers
The local organising committee consists of the following members:
The scientific committee consists of the following members: